Previous experiments are based on the data generated by the file gendata_full.m in the LSCS_KFCS_code.zip. I noted that the data actually are not stable, since the data is generated by the command:
x(:,t) = x(:,t-1) + sqrt(Q1)*randn(m,1);
So, the energy will increase with time. It is interesting to see when the the system x(:,t) is stable (i.e. X(:,t)=P X(:,t) + V, with the eigenvalues of P are less than 1), what's the performance of the algorithms. The stable system is more often encountered in practical problems.
So, I carried out another experiment using stable data. Each coefficient time series was generated as:
x(i,t) = b * x(i,t) + sqrt(1-b^2) * randn(1), with b randomly drawn from [0.7,1).
So, each x(i,:) is a temporally correlated time series.
At initial stage, the number of nonzero coefficients were 15. The support of x(:,t) will change at t=15, 25,30. New 10 nonzero coefficient time series were added in the support at t=15 and 30, while 4 existing coefficient time series were removed at t=25.
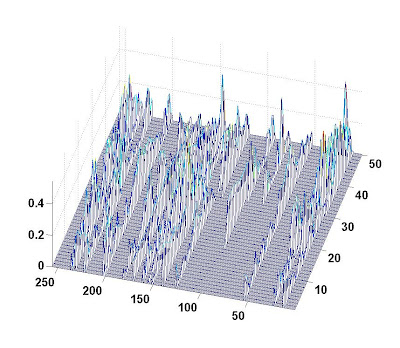
A picture of the evolution of coefficient amplitude is given below (the label for the bottom axis is the index of coefficient; the label for the right axis is the snapshot index; the label for the left axis is the absolute amplitude):
The dictionary matrix was 60 by 256. The total number of snapshot was 50. Noise standard variance was 0.01 (about 20 dB). I compared KF-CS, M-SBL and my T-MSBL. KF-CS was performed on the whole data. T-MSBL and M-SBL was performed on 5 segments (I evenly divided the whole data into 5 short segments with 10 snapshots). Experiment was repeated for 25 trials. Here is the result:
We can see T-MSBL achieved the best performance. I am somewhat surprised by the behavior of KF-CS. Probably KF-CS's detection to disappearing coefficients is more sensitive to the pre-defined values. As for elapse time, KF-CS cost about 190 second, while T-MSBL and M-SBL cost about 3 seconds (note here the input argument MIN_GAMMA for the SBL algorithms was set to 1e-3, which was different to the values in previous experiments. Different values of this input argument can result in significant change in speed).
The strange behavior of KF-CS in above experiment is probably due to the algorithm sensitive to the pre-defined threshold to remove disappearing coefficients. I used the threshold value given by the code, but didn't try other various values, because each running took much time. So, to remove the effect of this issue, I designed another experiment.
The experiment is an exactly MMV experiment, i.e. the support of each column in X didn't change (all satisfy the common sparsity assumption). Here is the result:
As in the above experiment, T-MSBL and MSBL were performed on 5 segments. And their final results were the concatenation of the recovered results from each segment. Again, we obtain the same conclusion.
So, I believe, for time-varying sparsity problems, dividing the whole data into several short segments and then using MMV algorithms on each segment is an effective method.
Referece:
Z. Zhang, B.D.Rao, Exploiting Correlation in Sparse Signal Recovery Problems: Multiple Measurement Vectors, Block Sparsity, and Time-Varying Sparsity, ICML 2011 Workshop on Structured Sparsity
-----
Another tulip is flowering in my patio:



Zhilin,
ReplyDeleteHave you tried a similar experiment with modified-CS (http://home.engineering.iastate.edu/~namrata/research/SequentialCS.html ) ?
Cheers,
Igor.
Hi, Igor,
ReplyDeleteI didn't do the comparison. I'll find time to do such comparison. The results should be interesting. Once I've done, I'll tell you and post in my blog.
Thanks for the suggestion.
Zhilin